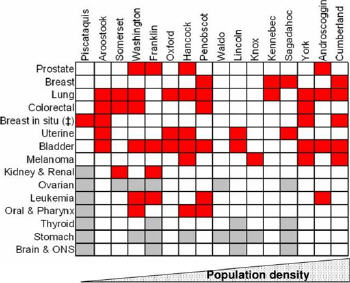
The obvious question is what combination of genetic, cultural and environmental factors is behind this unacceptably high incidence rate. The Maine Institute for Human Genetics and Health (MIHGH), a not-for-profit research subsidiary of Eastern Maine Healthcare Systems, has teamed with James W. Sewall Company (Sewall), a private Maine consultancy with specializations in mapping and GIS, to develop the Maine BioGeoBank, a research resource that links a repository of annotated human cancer biospecimens and cancer registry data with a GIS repository of cultural and environmental data. The BioGeoBank will enable researchers to undertake complex queries and analyses that explore relations between cancer genomics and the rural environment, yielding a better understanding about human susceptibility to cancer.
Why Maine?
Maine is a particularly useful test ground for this research. Our population is remarkably stable, and Maine's rural families tend to be large, extended and multi-generational, often living in close proximity to one another with similar lifestyles and environmental experiences. In Aroostook County, for example, 16-20% of householders have lived in the same residence for at least 30 years, compared to 10% in the U.S. overall. To cite another statistic, more than 70% of European families trace their genealogy to settlers of the 1750s. These sorts of factors enable better mapping of family genomics and histories than is often available elsewhere.
Maine's environmental history includes substantial use of toxins in ship building, forestry and the pulp and paper industries, as well as herbicides and pesticides used in Maine's blueberry and potato industries. Furthermore, an unusual geologic feature in Maine is the geographic juxtaposition of radon with endocrine-disruptive chemicals (EDCs) such as arsenic and dioxin. Arsenic occurs naturally in soil and bedrock and in particular is present at high levels throughout the area, as well as being used in certain wood products and as an ingredient in pesticides.
The BioGeoBank
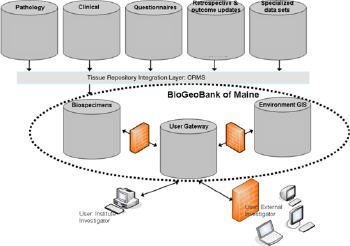 |
A user gateway enables complex queries drawing on data from the CRMS with the GIS-enabled environmental data. The GIS component is a relatively straightforward geospatial data management and delivery system. Sewall hosts the data in a PostgreSQL/PostGIS environment delivered using its GeoPower Hosted Portal Solution. Current data layers for the pilot areas currently under investigation include: streams, rivers and lakes for arsenic concentrations; geologic formations for radon emissions; soils for mercury; industrial use patterns for pesticide and herbicide applications; air quality; and wastewater outflows, brownfields and other EPA regulated sites. Each dataset is layered on appropriate land bases. More data layers will be added as they become available. The process is, of course, iterative. As more data become available, better correlations will emerge, calling in turn for new and better data. Security firewalls allow for levels of access to safeguard protected information.
Geospatial Challenges
There are significant practical challenges in using GIS data as envisioned here. One challenge, for example, is what is known in the GIS industry as the "quilt" problem. Many GIS datasets, especially those covering large, disparate regions, are assembled, or quilted together, from multiple sources, each with its own differing purpose and with its own scale, accuracy and reference points. Yet in such cases there are likely missing data that, because of their absence, will skew or bias queries.ï¿1⁄2 If one analysis matching cancer incidence with certain environmental data shows a high correlation of events in locations A and C, but not in a similar or intervening location B, possible explanations include the fact that the data do not support the correlation at B, or representative environmental or geospatial data at B equivalent to that at locations A and C were never captured. A related problem is that of trying to align and layer data captured with different scales and accuracies. What first appears to be a strong correlation between events and location-types may turn out completely different or even absent when the events are layered on a more accurate land base.
Another key geospatial challenge concerns the spatiotemporal requirements central to these studies. If, for example, the incubation period of a certain kind of cancer is 10-15 years and the research is querying correlations between that cancer and a certain type of chemical exposure, the subjects must be linked to particular locations where exposures were likely during the exposure period. Effective methods to incorporate the temporal dimension into GIS are only now emerging from conceptual and pilot studies.
Finally, few large GIS data acquisition projects have been driven by medical research questions. Projects that cover large geographic areas on a high resolution scale are often commissioned by public entities whose drivers are determined by political uses and boundaries, not medical or even environmental needs. Cancer registries, for example, capture patient data by political boundaries, with county often the smallest data aggregate when population density is low; yet exposure patterns linked to rivers, for example, typically transgress political boundaries on a regular basis. This problem speaks to increased coordination for data acquisition based on environmental and demographic patterns rather than political ones.
Conclusion
Given the complexity of issues surrounding the growth in cancer rates in Maine and throughout the nation, we believe the BioGeoBank's research potential is significant. The opportunities for GIS to contribute to this method of research are both exciting and challenging. The opportunities for discipline cross-fertilization are particularly exciting. We believe that clinicians' research needs will help drive development of effective temporal mapping, for example. Likewise, mappers' understanding of issues like the quilting problem should assist researchers in making better correlations and judgments. We expect the partnership of biomedical research and GIS to make meaningful contributions to advances in our understanding of human health.
____________
Acknowledgments: MIHGH is grateful for financial support of the U.S. Dept. of Defense grant number W81XWH-07-2-0116, PI: JM Hock, and Eastern Maine Healthcare System (EMHS). We would like to acknowledge Paul Laub, PhD, MIHGH BioGeoBank for his work on some of the descriptive epidemiology mentioned in the text.
Ed. note: A version of this material was originally presented at the URISA GIS in Public Health Conference, June 6-8, 2009, Providence, RI.